Plamu

Salmo salar (or “Atlantic Salmon”, “Black Salmon”, or “Plamu” as it’s known in my neck of the woods) is a culturally, ecologically, and economically vital fish species that has experienced widespread declines over the last century. Damming, habitat degradation, climate change, and aquaculture are all thought to pose significant threats to salmon health, but without consistent historical monitoring data, it can be difficult for scientists to determine which factors (and which interactions between factors) are contributing the most to this dwindling resource.
Slippery Monitoring
Managing fish as a resource requires knowledge of their life cycle, behaviour, and population trends over time. Population size is a particularly useful metric for measuring the health of a resource species. After all, how can you know what number of fish is ok to responsibly harvest if you don’t know how many fish are out there? In ecological terms, a “population” is a group of organisms who can reasonably be expected to reproduce with each other. Distinct populations may be separated by physical, temporal, or behavioural barriers that prevent individuals of different groups from mixing. A “metapopulation” takes this one step further and refers to a collection of multiple populations.
In salmon, populations are often isolated by strict fidelity to a particular spawning site, so that salmon from one river are unlikely to breed with salmon from another river. Together though, each population from every river draining into the Atlantic Ocean make up the Atlantic Salmon metapopulation. When considering the state of the species, we have to consider the larger Atlantic Salmon metapopulation, and this makes monitoring trends over time very complicated.
Surveys of fish populations can take many forms. Physical counts of captured individuals can be useful, but are time consuming, labour and resource intensive, and require a very large spatial effort to make sure the sample is representative of the whole population. In addition to these barriers, scientists must monitor populations for years to understand trends over time and determine causes of any witnessed declines.
Increasingly, genetic technology is being used to fill in the temporal gaps left by sparse historical monitoring.
Why You Shouldn’t Marry Your Cousin
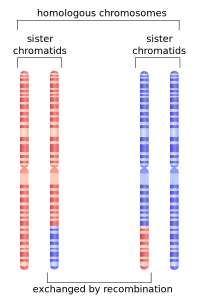
In salmon, humans, and just about every other thing on this planet that we consider to be “alive”, the information that controls who we are, is contained within these tiny chemical strands called DNA. DNA is arranged into chromosomes, which are usually found in these tightly-bundled packages inside the nucleus of our bodies’ cells (they kind of look like ramen noodles). During cell division, these chromosomes pair up in preparation for replicating and, sometimes, they get stuck together and end up switching or exchanging bits of themselves with the other chromosome (as if you cut two ramen noodles and then stitched the cut parts onto the opposite noodle’s end). Scientists call this process Recombination, and it explains why I look so much more like my grandmother than I do my grandfather, even though on average I should’ve received an equal 25% of my DNA from each of them. It also allows us to look into the evolutionary past of the organisms experiencing it.
Recombination happens at somewhat predicable but different rates, depending on the species and the location it occurs in the genome. Most of our DNA (and the DNA of salmon) doesn’t “code” for anything important, so bits of DNA getting recombined and moved around each time a cell divides usually isn’t a big deal. In a large population, you would expect to see a high degree of genetic diversity in these areas of the genome, and likely also in other areas that do “code” for traits but that aren’t vital for an individual’s survival. As a population gets smaller, however, and individuals are forced to reproduce with others more and more genetically similar to them, we see less diversity in these recombined areas. There are just fewer assemblage possibilities out there.
That can be really bad if the DNA getting passed around codes for something harmful, like smaller size, poor body form, or deadly heritable diseases. Low genetic diversity also makes small populations less capable of adapting to environmental changes, because they don’t have the same reserve of novel advantageous traits to fall back on that large diverse populations do. Scientists call this “Genetic Drift”, and it can wipe out small in-bred populations.
Piecing together the past
Knowing what genes code for different advantageous and disadvantageous traits, and what typical recombination rates look like in a particular area of the genome, allows scientists to both estimate past population sizes and postulate what environmental factors might be responsible for population declines.
Lehnert et. al., in a collaborative study conducted by researchers in Canada, Norway, Scotland, and the United States, published their attempt at reconstructing past population size of the Atlantic Salmon in Nature Communications earlier this year. Comparing DNA at linked and recombined sites in salmon from hundreds of populations, and then comparing their findings to survey data, they reconstructed population size for 172 populations across the entire Atlantic salmon range. What they found confirmed previous fears about the state of the species as a whole.
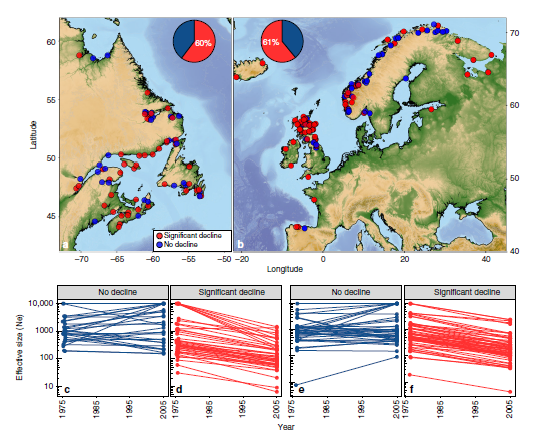
Recombination rates reveal population declines
Lehnert et. al. found that 60% of North American and 61% of European Atlantic Salmon populations have experienced significant declines in population size since 1975. These declines were particularly extreme in North American populations, with some falling to just 10% of past numbers. In both continents, most studied populations were estimated to contain under 1,000 individuals.
Climate changes and habitat disturbance explain most salmon declines
Facing a unique suite of threats in each river, the salmon populations showed a considerable degree of variation in whether or how much they declined over the study’s 30 year reconstruction. In North America, this variation was best explained by climate, particularly warmer winter temperatures, and human density, which the researchers considered to be a proxy for habitat disturbance. In Europe, it was all climate – especially climate variability/isothermality, warmer winters, and greater precipitation. However, the researchers note that, at this global scale, it’s likely that regional or local influences are being underemphasized, such as the interactive effects of a local dam development, small scale pollution, etc.
Declining and stable populations are evolving differently
If that news wasn’t already depressing enough, the researchers also found that these population declines are affecting the way salmon evolve. In both continents, salmon from declining populations were more likely to present identical DNA snippets in areas of the genome important for head formation, mucous secretion (mucous protects fish from external pathogens), migratory behaviour, and immune pathways. Should something happen, such as an environmental stressor that requires increased mucous production, or a particular immune response, these declining populations may be less likely to adapt to and survive under changing environmental conditions.
Atlantic salmon continue to decline across their range despite conservation efforts, with climate change being a major factor at the broad scale of this study and human impacts on habitat being especially important in North America. This study shows for the first time that there are genomic changes associated with declining populations of Atlantic Salmon across their range. As population size continues to decrease, genetic drift may come to play a larger role in salmon evolution than selective adaptation, making it harder for smaller and smaller populations to adapt to stressors like climate change. Are we witnessing the slow extinction of the Plamu?
If you’d like to help support Atlantic salmon research and conservation efforts, you might check out the Canadian Atlantic Salmon Federation, Mi’kmaw Conservation Group, Unama’ki Institute of Natural Resources, or the UK’s North Atlantic Salmon Conservation Organization.
Lehnert et. al. (2019) Genomic signatures and correlates of widespread population declines in salmon. Nature Communications, 10. | https://doi.org/10.1038/s41467-019-10972
Hi! I’m Rebecca Parker. I’m an ecologist and plant lover working in non-profit conservation in Nova Scotia Canada. I trained at Dalhousie and Ryerson University, where I completed a Masters in Environmental Science and Management. I like botany, wetlands, and wetland botany! On the sciencey side, I like to write about current topics in population and community ecology, but I’m also really interested in environmental outreach, how exposure to science and demographics affect environmental values and behaviours, and best practices for building community capacity in environmental stewardship. Check out my instagram for photos of the awesome nature I see through my work.
