Microbes can be found almost anywhere you’d imagine – in your own gut, on your cat’s fur, in the soil – even in the air you breathe. Some of these microscopic organisms can help maintain your health or the environment around you, while others can infect them with disease. Some provide nutrients to carbon-absorbing plants, while others emit greenhouse gases, such as carbon dioxide or methane, into the atmosphere.
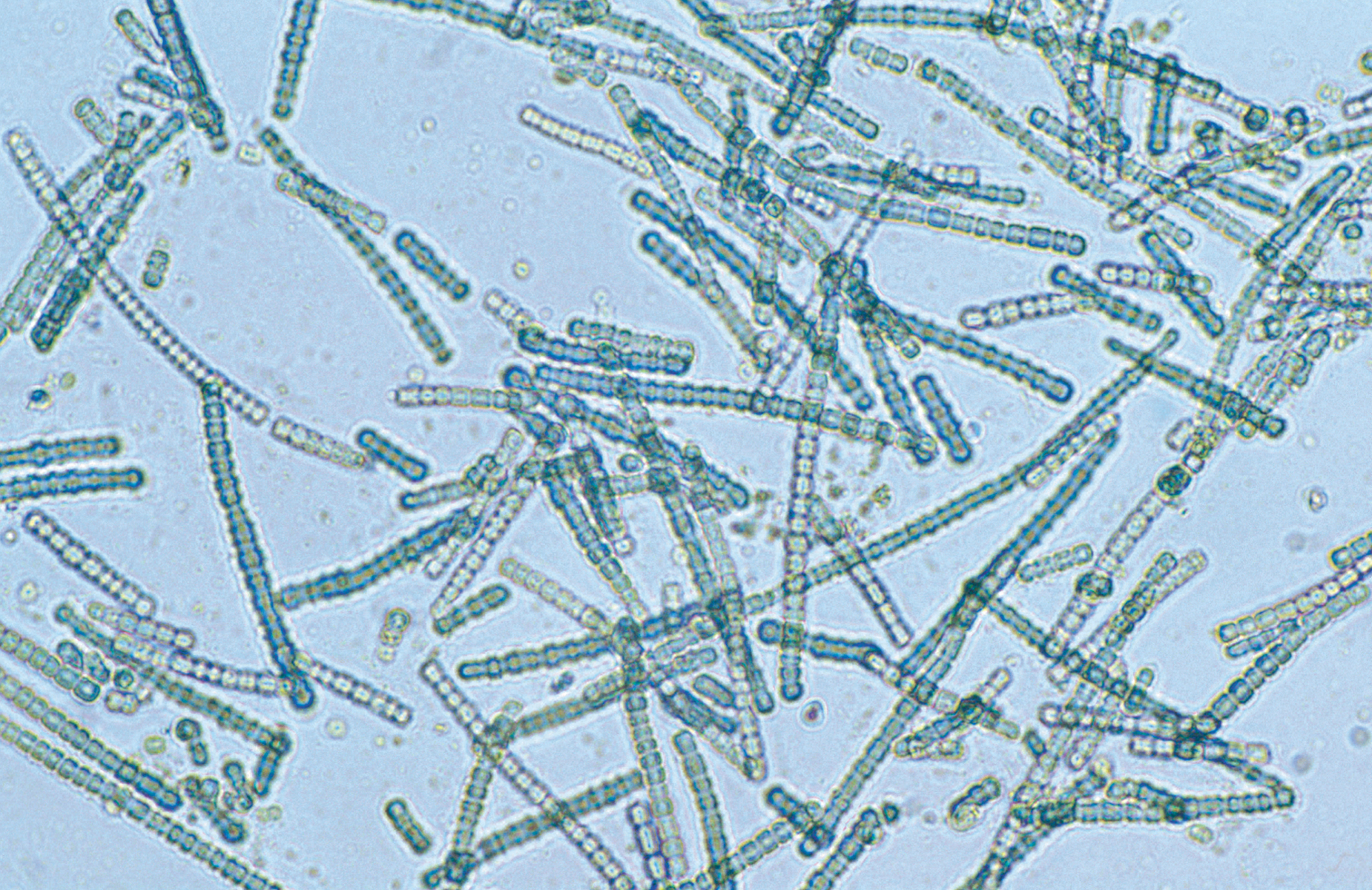
Because they can be so abundant, microbes are often studied in terms of their community composition – that is, how many species are present in a small sample? Which are the most abundant? How does one community compare to another? What functions do these communities have? To answer these questions, many scientists sequence the DNA of samples from particular environments they wish to study. The sequencing of DNA directly from an environment, known as metagenomics, involves decoding all the genetic material contained within an environmental sample to collect information about it, such as what kinds of species it contains.
As more is revealed about the roles microbes play in different habitats, scientists hope to use metagenomics to learn how these roles might change as the environment does. For example, lakes are known to change with (or in response to) climate, and may be a significant contributor to methane emissions – a greenhouse gas much more potent than carbon dioxide. In a recent study, researchers aimed to determine whether small- or large-scale drivers – for example, nutrient availability in lake sediment versus overall biological productivity of a lake – drive the evolution of microbial communities there.
Creatures at the bottom of the lake
Three types of microbes were monitored – bacteria, archaea, and fungi – each differing in cellular structure and energy transformation. To vary the large-scale environment, the researchers chose two distinct, relatively undisturbed Canadian lakes as their test sites. Swan Lake and Lake Laurentian represent two trophic statuses – oligotrophic and mixotrophic, respectively. The first contains much fewer nutrients, and is thus limited in its habitability for microbes, who rely on nutrients as their food. The second contains slightly more nutrients, and is thus more favorable to microbial growth.
To vary the small-scale environment – resource (or nutrient) availability – the researchers set out identical sets of mesocosms at the bottom of each lake near shore. Mesocosms are controlled tests using isolated containers set up within the natural environment – a mix between a field and lab experiment. In this case, the mesocosm set-up was designed to observe the impacts of differing types and quantities of organic matter (OM) within the sediment – a nutrient-rich by-product of plants and animals consisting mainly of carbon that microbes like to feed on, such as old leaves or feces.
Each site contained three different fractions of OM in the sediment – 5%, 25%, and 50% – for three different mixes of OM. The mixes contained leaves, branches, pinecones – any organic material that had fallen to the ground in forests nearby. One contained mainly deciduous (seasonal) plant matter, one contained mainly coniferous (typically evergreen) plant matter, and the last contained an equal mix of both.
After the OM was reduced to small particles, mixed with sediment, and placed at the sites, the researchers returned to the sites once a month from August to October 2015, and again a year later, in August 2016. Each time, they extracted pore water – water contained within the soil – from each mesocosm, and sequenced the DNA of each sample.
Microbes mainly respond to small-scale changes
The data revealed that bacteria and especially archaea quickly became distinct from their original communities after only two months, while fungal communities did not change at all. The most important driver of changes for bacteria and archaea was the fraction of OM in the sediment, representing the availability of microbe food in the small-scale. For both bacterial and archaeal communities, the mesocosm with 50% OM – the most abundant amount – diverged fastest in terms of species make-up.
It is common for the species make-up in a microbial community to change along with the genes that enable certain functions, but occasionally, certain dominant genes will remain the same as the species make-up continues to evolve. This ecological concept is known as functional redundancy. Thus, the researchers also tracked specific genes associated with functions microbes carry out – such as the transformation of nitrogen or production of methane. A year after the mesocosms were set up, the genes associated with the decomposition of OM – such as those that produce methane – were found to increase regardless of the altered species make-up. This means that although the communities looked much different over time in terms of species make-up, they still converged on existing roles (or functions) in their environment.
Global effects of the microscopic

The implications of these results are far-reaching. First, as carbon emissions from human activities continue unabated, these results imply that lake bacteria and archaea will continue to change at an accelerated rate as more organic matter (carbon) is buried at the bottom of lakes. Second, and more concerning, is the fact that these changes may be accompanied by an increase in methane-producing microbes – methanogens – which could increase the already significant global methane emissions of lakes (6-16%) and contribute even more to the greenhouse gas emissions driving our climate crisis.
As both small- and large-scale conditions continue to change, studies like these will be necessary to determine the many roles microbes play in our world, and how these roles will evolve to suit the environments we create – for better or worse.
I’m a PhD student at the University of Rhode Island’s Graduate School of Oceanography. I use a small-scale computer model to study how physical features like surface waves at the air-sea interface produce friction for the wind that can limit momentum, energy, gas, and heat exchange between the ocean and atmosphere. In the future, I hope to learn more about the role waves play in different parts of the world as weather and climate patterns evolve. Also, I love to write.


