Reference: Needham, D.M., Yoshizawa, S., Hosaka, T., Poirier, C., Choi, C.J., Hehenberger, E., Irwin, N.A., Wilken, S., Yung, C.M., Bachy, C. and Kurihara, R., 2019. A distinct lineage of giant viruses brings a rhodopsin photosystem to unicellular marine predators. Proceedings of the National Academy of Sciences, 116(41), pp.20574-20583.
DOI: 10.1073/pnas.1907517116
What are marine viruses and why are they important?
Viruses are found across the globe and can thrive in a variety of different ecosystems including terrestrial, freshwater, and marine. In fact, some viruses can even be found in deep ocean sediment. Unlike many terrestrial and freshwater viruses, marine viruses are relatively understudied. Most known viruses infect species with human impact, such as harmful algal blooms that decimate fisheries and cause respiratory symptoms in humans. Viruses have recently been found to be an important part of biogeochemical cycling in global oceans as they break open cells, releasing a variety of nutrients and organic matter to be recycled back into the marine food web. In order to study viruses, methods still rely on techniques that require cells viruses infect (i.e. host cells) to be cultured in the laboratory. This presents a challenge for many organisms as both prokaryotes and eukaryotes are near impossible to grow in a lab without knowing what medium they can grow on (i.e. what food they like) and what environmental conditions are preferable. Many organisms cannot be cultured in lab which is why many researchers now rely on molecular techniques to identify and characterize unculturable marine prokaryotes and eukaryotes. Needham et al. (2019) set out to use cultivation independent techniques and newer laboratory methods to investigate eukaryotes and their associated microbiome (bacteria and viruses).
How to study the unculturable
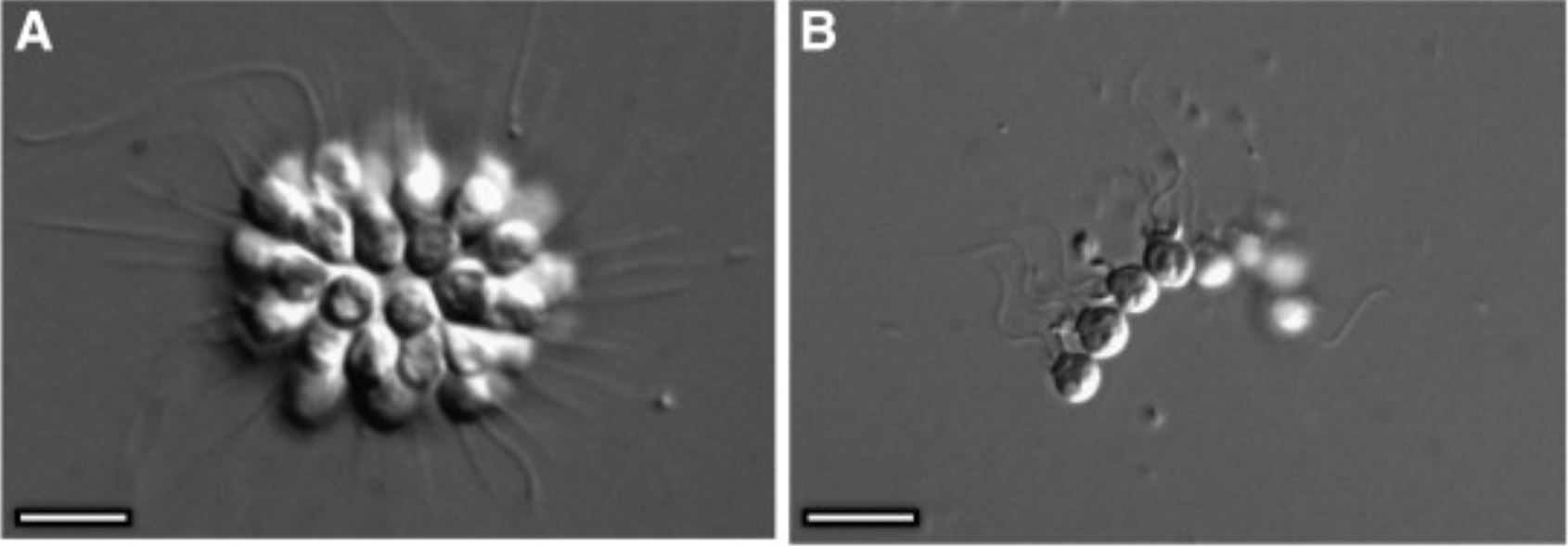
The authors collected water, filtered the water to exclude smaller eukaryotes, and stained the filters to use in an instrument that can sort individual cells. This instrument, known as a flow cytometer, uses different parameters such as fluorescence, or the wavelengths emitted when exposed to varying wavelengths of light. Fluorescence can be measured from stained cells or natural fluorescence from photosynthetic pigments in algal cells and used to separate and pick out cells of interest. This study focused on predatory eukaryotes and therefore specifically excluded these algal cells with a photosynthetic pigment. At the end of each run, a plate with ~ 300 wells was filled, one cell per well. These cells were then used for species ID using a marker gene, which uses a genomic region that is similar across all organisms of a particular group (in this case predatory eukaryotes) with enough variability to differentiate between different organisms down to genus or species. After the DNA sequences were analyzed, a species of predatory protist with a large virus was discovered. The authors then targeted this species, Bicosta minor, for single cell genomics. Single cell genomics is a molecular technique that involves extracting and sequencing all DNA available in an organism to discover the capabilities of each cell. Metatranscrimptomics of bulk RNA for these four B. minor cells was also conducted, and is a technique that uses RNA to reveal which viral genes are actively being used in each B. minor cell. Finally, the authors discovered an important viral gene coding for a protein that allows host cells to respond to light. In order to characterize this protein, experiments were done to investigate how the protein reacts to varying pH levels and it’s structure.
What did they find?
In this study a new virus was discovered in B. minor, a predatory protist under the Choanoflagellate group. This newly discovered virus (ChoanoV1) was termed a ‘Giant Virus’ because of the size of its genome –875,000 base pairs of nucleotides. Additionally, the authors compared the ChoanoV1 virus to other sites based on nucleotide content characteristics. This mean they compared the number of guanine and cytosine bases to the number of adenine and thymine bases. They found another Choanoflagellate virus that contains 89% of the same genes as the originally identified ChoanoV1 virus and termed this ChoanoV2. Both viruses were compared to previously identified viruses and were found to share only 50% of the genes coding for proteins. These findings highlight how important this viral discovery is and how many gaps there are in sequencing data for both cellular organisms and viruses.
Why care about the genes these viral genomes carry?
Of the genes found in the viral genome, many are common across previously discovered viruses, including genes for viral replication, and recombination and repair in the host. However, some genes from the viral genome were more surprising, like those for a chitinase enzyme and a unique rhodopsin. Chitinase is an enzyme that breaks down cellular walls made from chitin. The authors hypothesized this enzyme could be used to break apart cell walls during infection or may help with prey degradation in the host cell. If viruses are able to directly break down host cell walls, this can have big implications for small-scale cycling of nutrients.
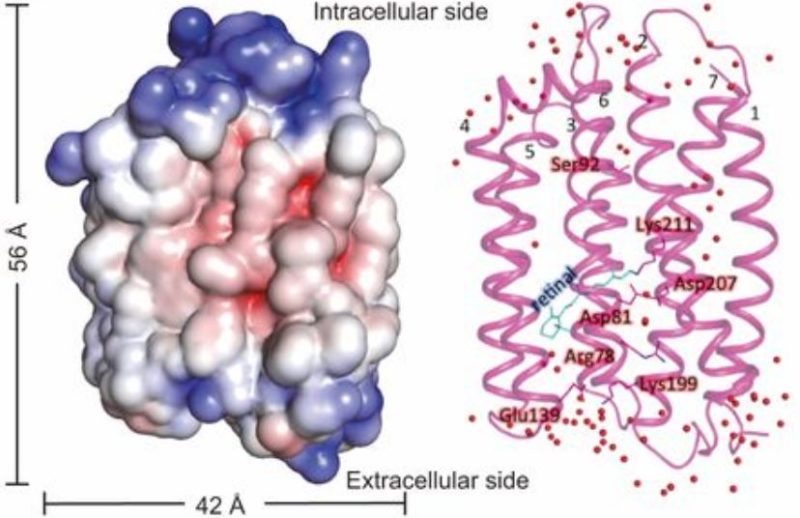
The rhodopsin gene found in the ChoanoV1 virus is unique and known in bacterial cells to allow a cellular response to light. These proteins are thought to help with cell motility and photosensitivity, by reacting to light. Some are known to be accompanied by a proton pump that allows cellular organisms to create energy using light. This process may sound familiar as photosynthesis uses a similar method, of a series of membrane-bound proteins to use light to create energy. After further experimentation with this rhodopsin protein, the 3-D structure of this gene was discovered as well as its response to varying pH gradients. Tests that show a pH response can indicate whether a proton pump is in use by the protein, very similar to the methods used in photosynthesis. The authors concluded that this rhodopsin protein likely did include a proton pump and because of this, could potentially alter where the host derives energy. This would be like only being able to get energy by eating food one day, but after a viral infection you can suddenly start getting energy by sitting in the sun. These viruses may use this method to alter the way the host can get energy and, in a way, alter this organism’s position in the food web.
What’s the big deal?
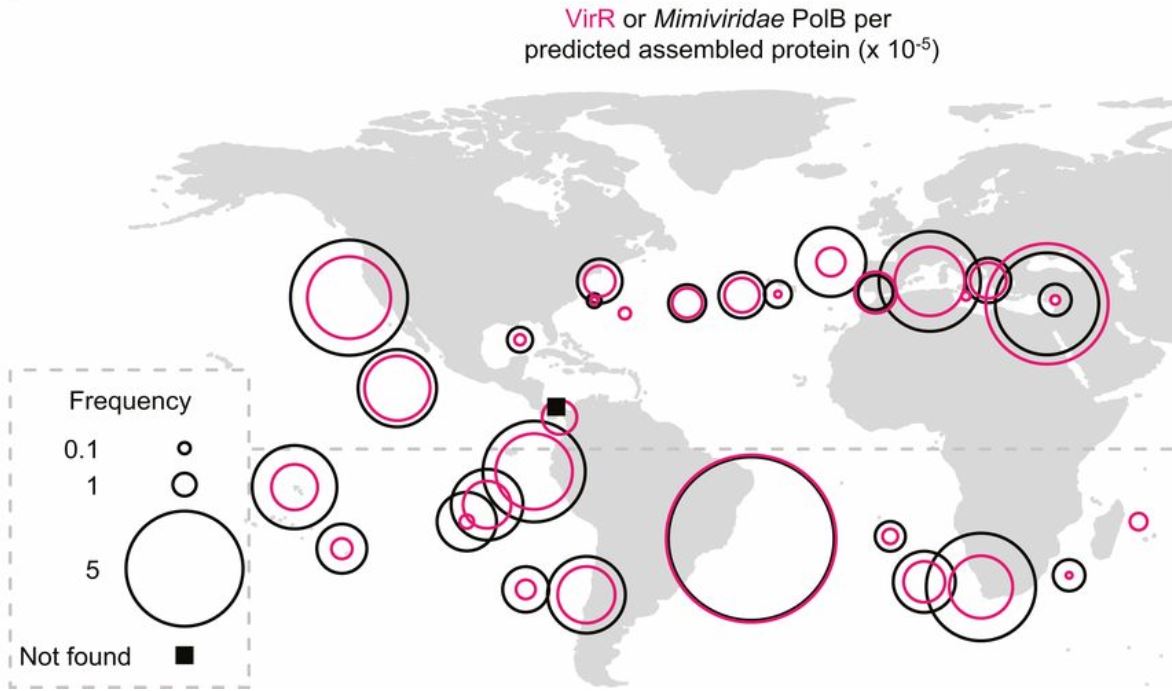
This study discovered the largest viral genome to date in the marine environments and further showed that this virus is ubiquitous across ocean regions. Not only this, but the authors discovered the viral genome encodes, and actively translates a rhodopsin gene that might be used to manipulate the way its host derives energy.
Directions for future research
Although these authors used a variety of new techniques in a creative way to reveal a new virus present in Choanoflagellates, they point out that there are still many gaps in research from marine viruses to the viral rhodopsin gene. They encourage more research on marine viruses and subsequent submission of viral genomes to increase their representation in databases. In addition, they recommend further study into the rhodopsin gene and its presence in global ocean waters. Finally, the authors mention a need for more studies regarding these co-associated entities, such as viruses, which can further reveal symbioses at play in nature.
Marine viruses are an interesting new horizon in our ongoing study of the world’s oceans and test the limits of our current knowledge.
Additional References
Dayel, M.J., Alegado, R.A., Fairclough, S.R., Levin, T.C., Nichols, S.A., McDonald, K. and King, N., 2011. Cell differentiation and morphogenesis in the colony-forming choanoflagellate Salpingoeca rosetta. Developmental biology, 357(1), pp.73-82.
I’m a PhD student in the Rynearson Lab at the University of Rhode Island (URI) Graduate School of Oceanography (GSO). My research interests are focused on human impacts on the oceanic ecosystem, particularly effects on the primary producers (phytoplankton) at the base of the food web. Currently, I work with cultures from regions of the ocean that are nutrient limited and will conduct experiments to investigate how these phytoplankton survive.


