Citation: Rey, A., Carney, K.J., Quinones, L.E., Pagenkopp Lohan, K.M., Ruiz, G.M., Basurko, O.C., Rodríguez-Ezpeleta, N., 2019. Environmental DNA Metabarcoding: A Promising Tool for Ballast Water Monitoring. Environ. Sci. Technol. 53, 11849–11859. https://doi.org/10.1021/acs.est.9b01855
Some background on ballast water regulations
Merchant shipping is estimated to transport about 3500 million tons of ballast water (BW) per year. Ballast water is taken up and discharged by ships as they transport goods around the world via our oceans. The water is used to maintain ship stability when it is not fully loaded with goods. BW provides an easy opportunity for thousands of species to be transported across different biogeographic regions. Imagine this: a ship travels from Japan to Los Angeles, unloads its cargo and then suddenly becomes much lighter than it was. To offset this missing weight, the ship takes in BW into its tanks for the trek back to Japan. Then once in Japan to get more materials, the ship offloads the BW to make more room for goods, unloading “Los Angeles” water into Japanese waters.

Do you see the problem? To combat the spread of invasive species throughout the world’s oceans, the International Maritime Organization has created regulations for cargo ships to follow. Currently, there are two main ways ships manage their BW: 1) they can exchange coastal water with oceanic water at a certain distance from major ports or 2) they can treat the BW using various technologies such as adding chlorine to the water, taking oxygen out of the water and manually taking organisms out through filtration. Monitoring the efficiency of these methods is important because it can help assess the potential “spread risk” of various organisms. However, it is also challenging because until recently, most BW surveys have relied on taxonomic expertise, meaning using a microscope to identify an organism by its small features. This method is limited because there are many different organisms found in BW and it is difficult for any one person to have the taxonomic expertise necessary for such identifications. Thus, scientists are turning to other methods for identification of organisms in BW. A recent study by Rey et al. uses genetic techniques to uncover the hidden diversity in ballast tanks.
So what is this eDNA method that the authors used?
Since taxonomic identification of organisms in BW can be difficult, there has been increased interest in using metabarcoding simultaneously with taxonomic assessment to compare results obtained from each method. Environmental DNA (eDNA) metabarcoding involves collecting DNA from environmental samples instead of directly from the organism. For example, this DNA is released into the environment and sampled from water or soil. This method allowed the scientists to simply collect water from the various parts of the ballast tanks and analyze the water with genetic techniques instead of having to deploy a plankton net. Analyzing BW with eDNA metabarcoding is a cost and time efficient method to uncover organismal diversity that may be missed by traditional morphological identifications. Often times, cargo ships are in port for a short amount of time and thus, a relatively quick water grab might be the only option scientists have to test BW. To the authors’ knowledge, their study is one of only a few to use the eDNA metabarcoding method for BW monitoring.
How did they do it?

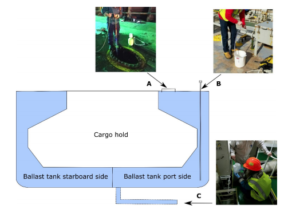
Rey et al. sampled water from 11 vessels entering Baltimore, MD and Norfolk, VA ports. They had multiple goals to offer insight about using eDNA metabarcoding as a tool for BW monitoring. These goals included: 1) whether different BW sampling access locations influenced the communities (See Figure XX for clarification), 2) whether the effect of common BW characteristics (i.e. how long the BW has been sitting in the tank, the type of BW management or the location of source) on communities could be identified and 3) whether eDNA could be used to identify taxa with comparisons to plankton samples identified with taxonomy. It is also important to note that the authors identified common taxa discharged frequently into the two ports to detect potential invasive species introductions.
What did the scientists find and what does this mean for the future of BW management?
Overall, there was more diversity observed with eDNA than by morphological identification. All groups of organisms identified by morphology using a microscope were also found with eDNA, except for a sample from one vessel. The age of BW in a tank was an important factor for community diversity—younger BW was more diverse than older BW. In one of the vessels, different sampling access locations from the same tank provided different communities at a higher taxonomic level. In addition, samples from the sounding pipe were less diverse than those retrieved from either the manhole or the discharge line. For BW monitoring, the sounding pipe can be easier to access than the manhole. However, the results of this study indicate that lower diversity is recovered from the sounding pipe so future BW management techniques should take note. If only the sounding pipe is sampled, there may be non-native organisms that are not accounted for.

The authors suggest that BW testing use in-line sampling through the discharge line because in-tank sampling could be affected by the tendency of organisms to exist in patches, meaning they may not be evenly spread out within the tank; there could be more organisms in one region over another. BW exchange will eventually be phased out as a management method and replaced with treatment systems to meet IMO standards, such as ships only allowed to have a certain number of living cells/mL in their ballast water upon discharge. Therefore, method development for assessing the efficiency of BW treatment systems is quickly increasing. The authors indicate that eDNA metabarcoding can be used to identify BW communities and differentiate diversity between tank sampling locations. It is also possible that eDNA metabarcoding can detect benthic organisms present in BW sediment through the release of these organisms’ DNA into BW. Traditional taxonomic methods would be unable to identify benthic organisms if they are not physically located within the sample. Using eDNA methods, the authors found eight species in vessels not previously found in the Chesapeake Bay. Thus, metabarcoding can be a useful method for detecting non-native species and limiting their expansion by helping to inform BW managers. The authors do note, however, that care should be taken with the identification of new Chesapeake Bay species using this eDNA technique because these species were identified with a particular genetic marker that can have poor taxonomic resolution. Thus, one of the major limitations in using eDNA methods for BW management is the lack of complete databases for DNA sequence matching to species. Despite this, the research shows that as more eDNA studies are completed with simultaneous taxonomic identification, databases will become more complete, enabling more thorough species identifications in ballast water and reducing the spread of non-native organisms.
I love writing of all kinds. As a PhD student at the Graduate School of Oceanography (URI), I use using genetic techniques to study phytoplankton diversity. I am interested in understanding how environmental stressors associated with climate change affect phytoplankton community dynamics and thus, overall ecosystem function. Prior to graduate school, I spent two years as a plankton analyst in the Marine Invasions Lab at the Smithsonian Environmental Research Center (SERC) studying phytoplankton in ballast water of cargo ships and gaining experience with phytoplankton taxonomy and culturing techniques. In my free time I enjoy making my own pottery and hiking in the White Mountains (NH).

